Important Details
Name : Backend Website Student Project for BioJS
Organisation : Open Bioinformatics Foundation
Mentors: Dennis Schwartz, Rowland Mosbergen, Yo Yehudi
Proposal : Project Proposal
Project Description (by org) : https://obf.github.io
GitHub repository (backend source code) : https://github.com/biojs/biojs-backend
GitHub repository (Ansible playbook) : https://github.com/biojs/biojs-backend-ansible
Live Website : http://43.240.98.213/
Overview
BioJS registry is a platform used by bioinformaticians and researchers all over the world. It serves as a common ground to index all the BioJS components (eg: Cytoscape) and their visualizations. BioJS had two websites, biojs.io (the registry) and biojs.net (an educational portal having guides). The goal of the project was to re-engineer the registry to render data and visualizations in minimum time possible to enhance user experience of the international audience. The website was a bifurcated into two projects, the Frontend project, which was achieved using VueJS, and the Backend project, which was developed using Django, a Python framework.
Summary
Project Goals
- The primary goal of the project was to either re-engineer or optimize the current engineering of the biojs.io registry, while prioritizing the improvement in the loading time of the same.
- In order to improve the user experience as well as the user interface, the registry website was designed from scratch, and APIs for the same had to be written and integrated.
- Write tests to ensure proficient working of the new components, especially the visualizations.
- Write an Ansible playbook to facilitate quick and automated deployment of the website in the future.
Goals Achieved
- A new workflow was created for the new website, along with a re-engineered mechanism that would enable quicker display of visualizations, an important part of the original repository.
- A new backend based on the proposed model as well as necessary changes made in accordance with the community decisions was made and the same was inegrated in unison with the frontend project.
- The django-ansible stack was created in the required time, and the same was tested across Ubuntu images pertaining to different versions.
- After intense discussions, a cron job was implemented to update the database timely from the npm registry as well as Github. The data requested via the API calls was returned after efficient querying from the database, rather than natively through Github and/or NPM registry. This increased the speed drastically.
- The data now loads at least 4 times faster than that in the previous website.
A peak of the new website

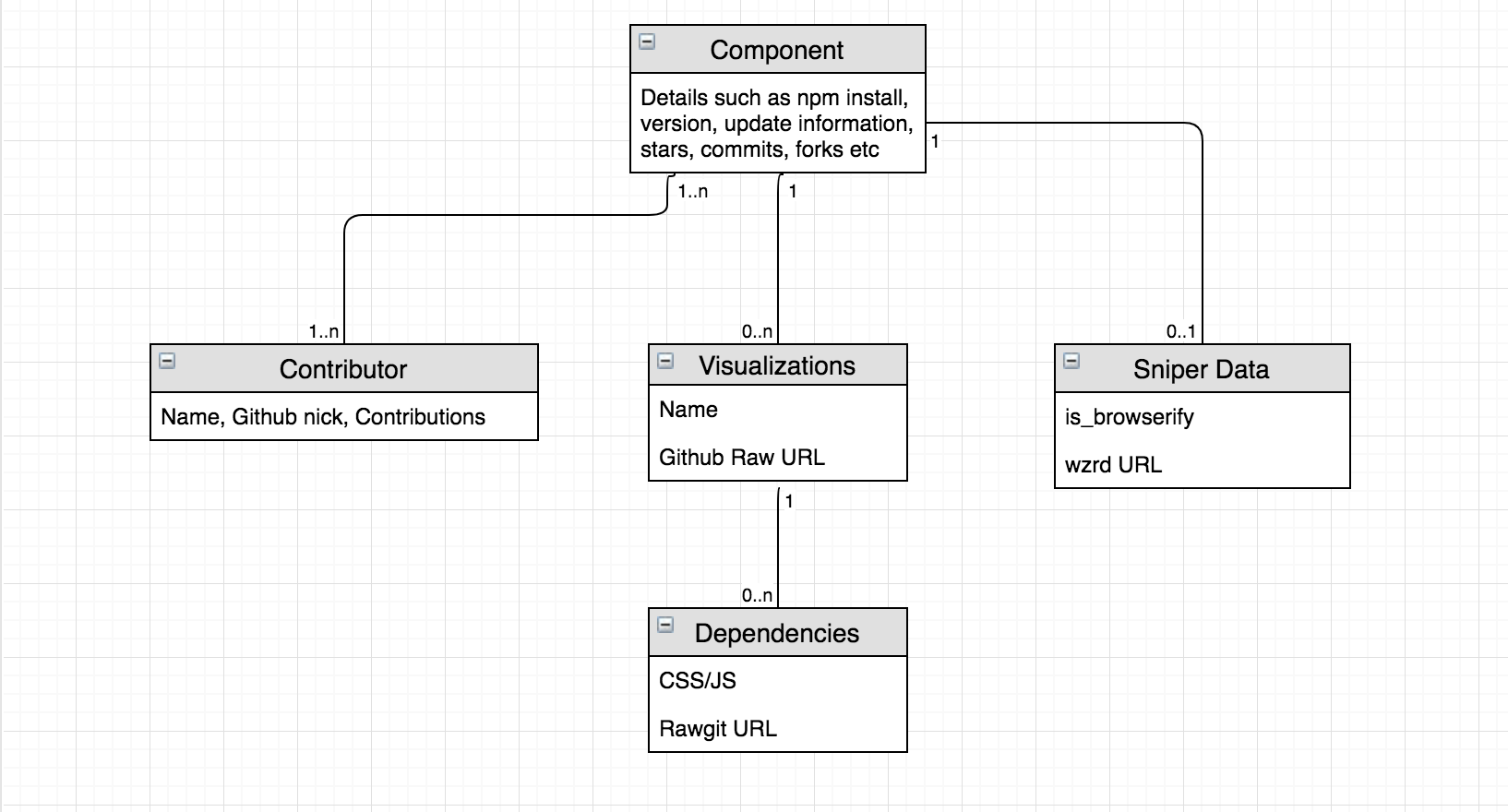
A class diagram of the database models used
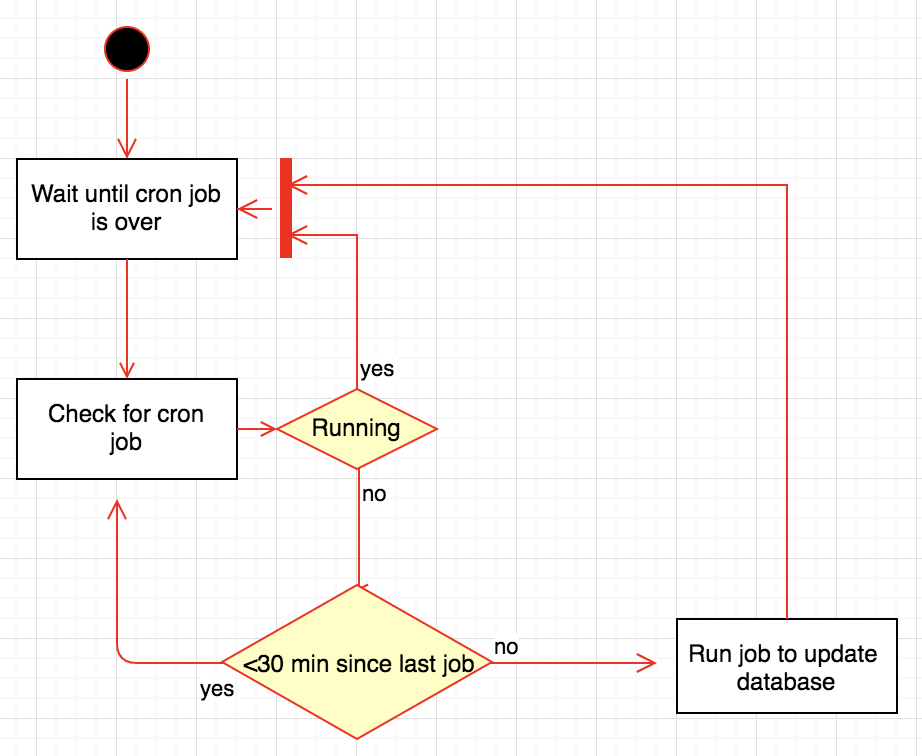
A very basic idea of the workflow implemented
Tech Stack Used
- Python
- Django
- PostgreSQL
- Github API
- TravisCI
- Ansible
- Gunicorn
- Nginx
- crontab
Coding Period Experience
It was an amazing experience, working with an international community, of developers, researchers as well as administrators. Apart from the technical aspects, I learnt a lot about important practices that need to be incorporated when working in a team, especially the correct way of communication and its importance. I am heartly thankful to my mentors for helping me not just improve my coding skills, but also skills that would help me in every step of my life.
There were weekly(not limited though!) video calls with my mentors, where we talked about tasks completed in the week, problems faced, their solutions, future tasks as well as improved scope for the project. Apart from these, we would communicate timely whenever we faced a problem or had to carry out tests that would request the help of my mentors. They were extremely enthusiastic for the same. Though they started as mentors, I am sure I made some great friends across the globe!
My weekly updates can be seen here.
Future Work
The following tasks can be considered for the future development of the project –
- Find a way out to host the visualizations without having to use wzrd as it is unstable and not something BioJS would like to be dependent on.
- Automate some minor tasks while using the Ansible playbook.
- Optimize speed further using server side caching (Redis or memcached).
- Some further changes have been opened up as issues in the repository.